What if you could follow the evolution of AML to uncover biomarker insights?
The complexity of leukemia clonal architecture and the genotypic and immunophenotypic drifts during treatment explain why more than half of patients in remission could relapse. Using single-cell multiomics approaches, researchers can uncover the clonal and subclonal drivers of therapy resistance and relapse, and more precisely characterize patient tumor heterogeneity and evolution.
Key Benefits
Early delineation of subclones driving myeloid progression, therapy resistance, and relapse.
Single-cell characterization of SNVs, indels, CNVs, zygosity, and surface protein expression to unravel clonal heterogeneity.
More cost-effective and easier than combining bulk NGS and Flow, with sample multiplexing and automated data integration.
Primary Applications

Clonal Heterogeneity
- Disease characterization
- Mutation acquisition
- Clonal architecture and evolution from precursor to disease state

Therapy Resistance
- Understand mechanisms of resistance
- Differentiate non/responders
- Identify and detect rare relapse clones

Disease Monitoring
- Sensitive and specific measure of residual disease
- Identify gene/surface protein targets for future therapeutic interventions
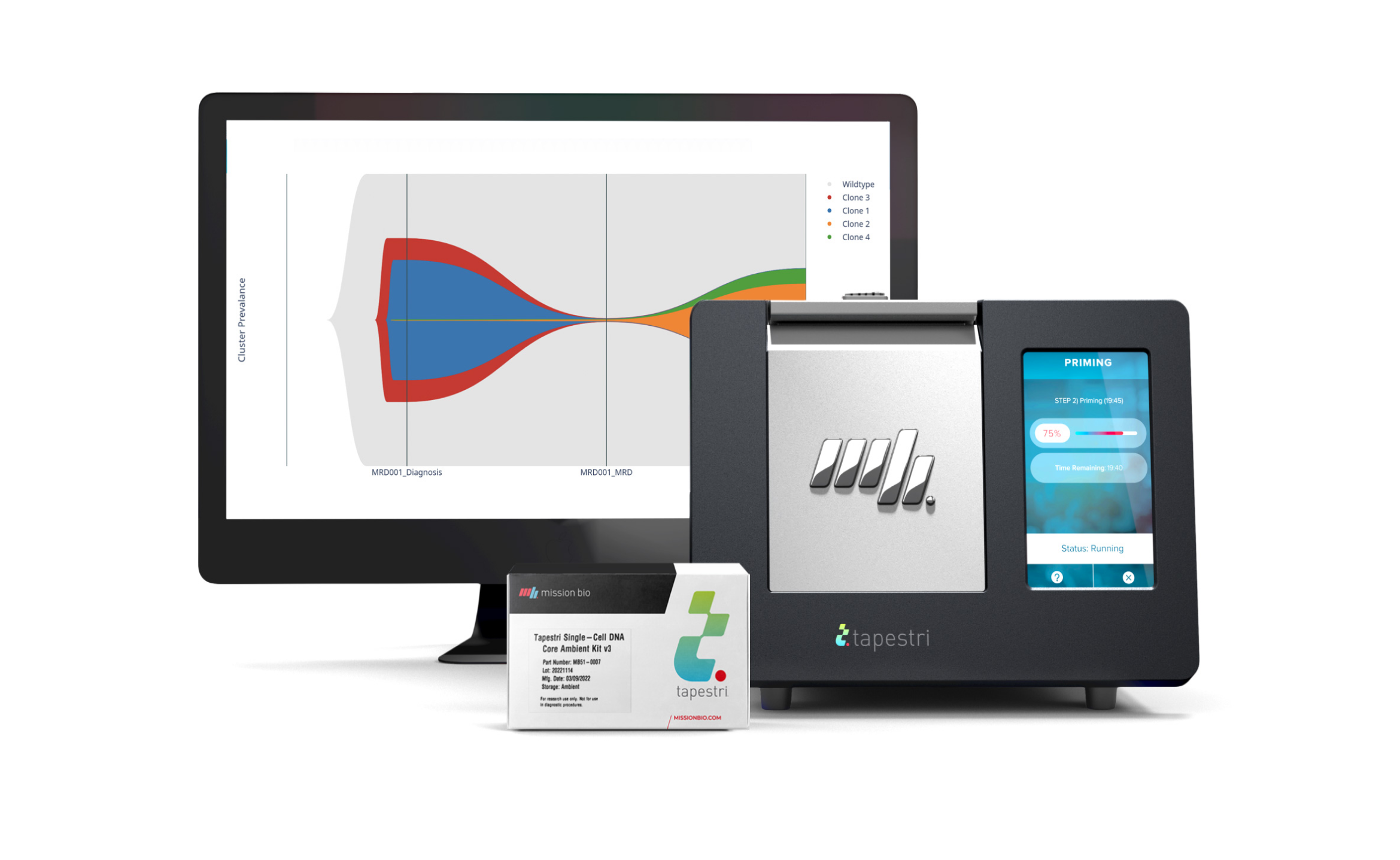
Tapestri Single-cell Myeloid Multiomics Solution
Get everything you need to conduct single-cell multiomics analysis of AML and other myeloid neoplasms in your lab with this fully-supported, all-in-one package.
- Tapestri Instrument
- Tapestri Single-cell DNA + Protein Core Kit
- Choose from Our DNA and AOC Panels for AML and Myeloid Diseases
- Software-Based Analysis (via Portal Access or On-Premise Activation)
- Sample Multiplexing by Genotyping
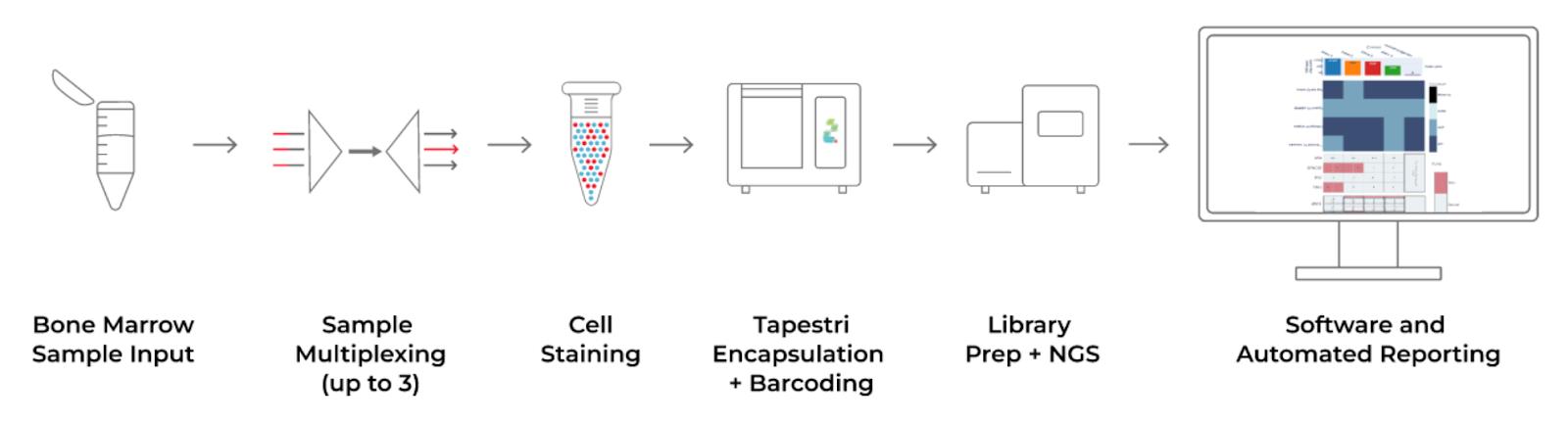
The Tapestri Myeloid Workflow
Flexible and Modular Assays
Panels can be purchased together or separately to suit your research needs.
| Panel | Targets Covered |
|---|---|
41 hotspot genes based on updated guidelines for acute myeloid leukemia progression, therapy resistance, and relapse. | |
37 hotspot genes commonly mutated in myeloid malignancies. | |
31 hotspot genes commonly mutated in myeloid malignancies providing insights into the pathogenesis of myeloid transformation. | |
17-plex antibody-oligonucleotide conjugate (AOC) panel curated for key biomarkers associated with AML. |

"Current tools for MRD assessment, like bulk NGS and flow cytometry, lack the clonal resolution and specificity to detect the treatment-resistant cancer cells still hiding in the shadows. Mission Bio’s unique approach to characterizing MRD could dramatically change how we stratify patients in clinical trials and create personalized care strategies in the future.”
P.J.M. (Peter) Valk , PhD, Principal Investigator at Erasmus MC
Independent Mission Bio Platform User
Resources

Molecular Analysis of Acute Myeloid Leukemia in Morphologic Remission
Get Started
Contact us to begin your evaluation.
For Research Use Only. Not for use in diagnostic procedures.