Track Cancer’s Most Critical Moments
Numerous genomic events and complex clonality in multiple myeloma (MM) render it a challenging disease with a high relapse rate despite advancements in therapies and diagnostic approaches.
Tapestri single-cell multiomics analysis transforms how you understand the complexities of multiple myeloma and its precursor stages by integrating genomic, proteomic, and clonotypic subclonal assessment with single-cell resolution, all within a single assay.
With the Tapestri Single-cell Multiple Myeloma Multiomics Solution, you gain crucial insights into the clonal and subclonal heterogeneity underlying myeloma progression, therapy response, and relapse often missed by traditional bulk methods, potentially improving the identification of novel biomarkers and the development of better therapies.

Hotspot Mutations in Driver & Resistance Genes
Assess clonal heterogeneity & evolution in multiple myeloma (includes SNV and focal CNA)

Zygosity
Discern zygosity of detected mutations impacting disease progression or drug resistance

Genome-wide CNV
Analyze whole genome copy number gains and losses

V(D)J Clonotyping
Identify Ig clonotype in the CDR3 region

Surface Immunophenotype
Classify cell types and identify immunotherapy targets such as BCMA and FcRH5
Primary Applications

From Stealth to Spotlight
SINGLE-CELL IDENTIFICATION OF SMM CLONES DRIVING MYELOMA PROGRESSION AND THERAPY RESISTANCE
Learn how to identify smoldering multiple myeloma (SMM) clonal populations progressing to active myeloma or resistant disease with a true multiomics platform that can quantitatively correlate multiple features from SMM subclones including SNVs, genome-wide CNVs, Ig clonotype, and surface protein expression.
Featuring Adam Sciambi, PhD
CTO, Mission Bio
Directly Correlate Molecular Signatures of Multiple Myeloma at the Clonal Level
Tapestri automatically generates comprehensive and intuitive reports with data visualizations to help you easily correlate various clonal and subclonal attributes.
Summary Map
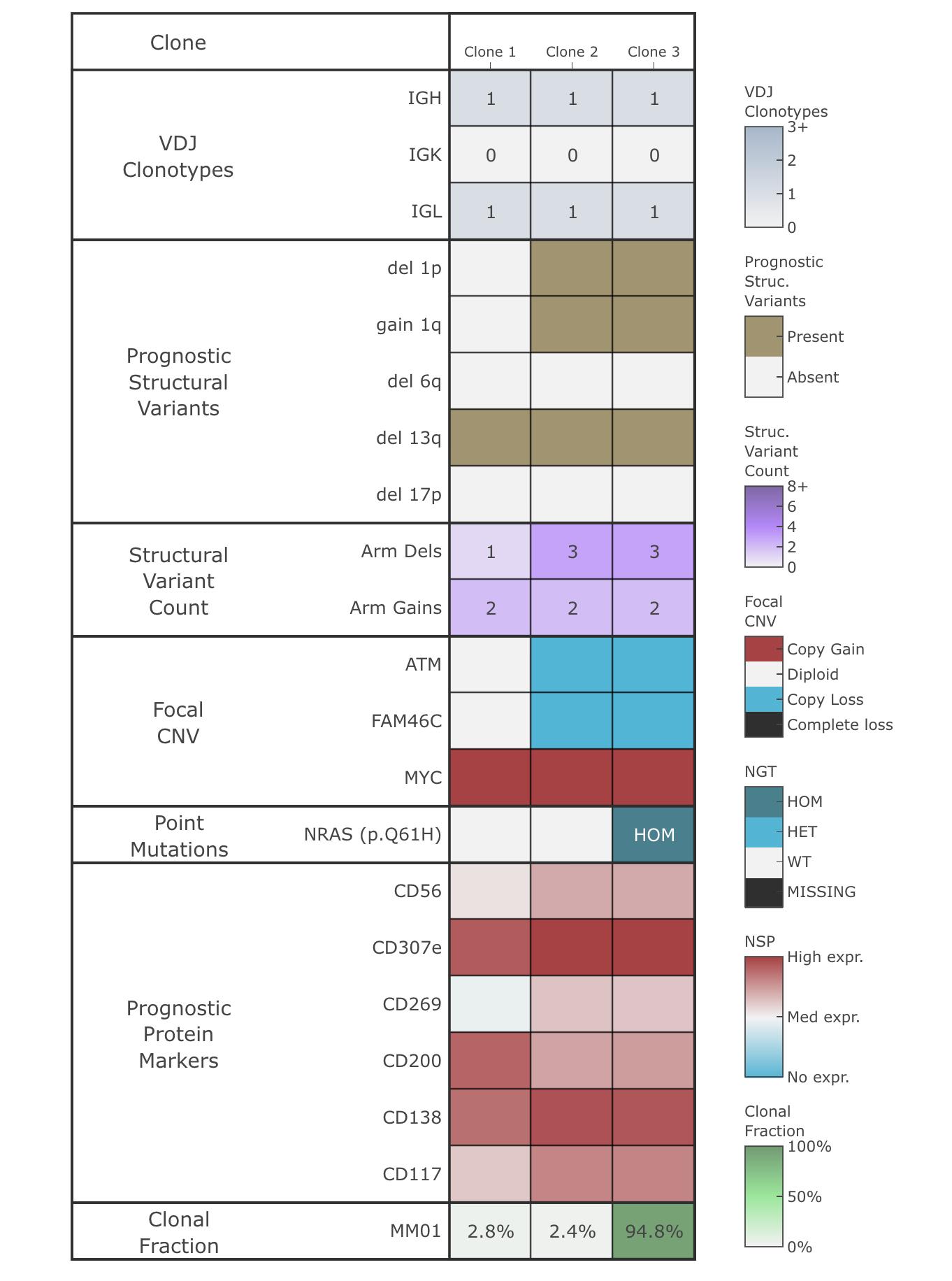
DNA and Protein Signatures

VDJ Clones and Proportions

Genome-wide CNV Profile

Protein Expression UMAP

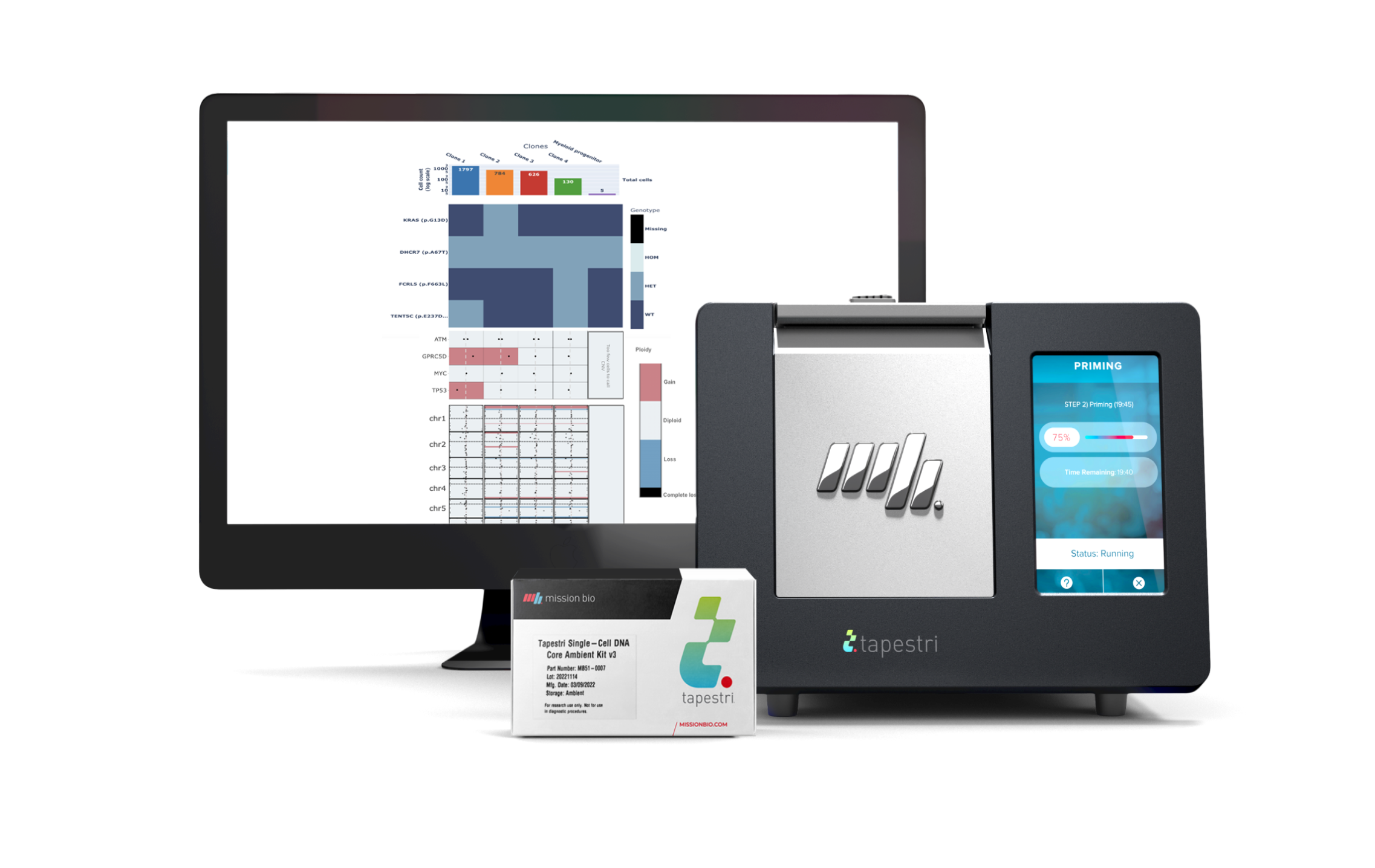
Tapestri Single-cell Myeloma Multiomics Solution
Get everything you need to conduct single-cell multiomics analysis of multiple myeloma samples in your lab with this fully-supported, all-in-one package.
- Tapestri Instrument
- Tapestri Single-cell Multiple Myeloma Multiomics Kit including:
- DNA + Protein Core Kit
- Multiple Myeloma DNA Panel
- Genome-Wide CNV Panel
- Multiple Myeloma Antibody Cocktail
- Software-Based Analysis (via Portal Access or On-Premise Activation)
- Sample Multiplexing by Genotyping
Assay Content
- Coverage of key driver and resistance genes, genome-wide copy number variations (CNVs), V(D)J clonotype, and cell lineage & immunotherapy markers
- Available to purchase as a complete solution or as standalone panels
Overview of Tapestri Single-Cell Multiple Myeloma Multiomics Assay Content
| Panel | Analyte(s) | Targets covered |
|---|---|---|
Tapestri Single-Cell DNA Multiple Myeloma Panel | Hotspot mutations in driver & resistance genes | 47 genes associated with multiple myeloma and therapy resistance |
Focal copy number aberrations (CNA) | 10 genes with focal copy number aberrations | |
V(D)J clonotype | CDR3 region of the IgH, IgK, and IgL chains | |
Tapestri Single-Cell DNA Genome-Wide CNV Panel | Genome-wide copy number variations | All chromosome arms except chrY |
TotalSeq-D Multiple Myeloma Antibody Cocktail | Cell surface antigens | 19-plex antibody oligonucleotide conjugates targeting cell lineage markers & therapeutic targets |
Details of Each Panel
34 genes with hotspot mutations
Drivers genes commonly associated with multiple myeloma curated based on
consultation with global experts and publications
ACTG1 | ATM | BRAF | CCND1 | CCND3 |
CDKN1B | CDKN2C | CYLD | DIS3 | EGR1 |
EZH2 | FAM46C | FGFR3 | IDH1 | IDH2 |
IKFZ1 | IRF4 | KRAS | LTB | MAF |
MAX | MYC | NFKBIA | NRAS | PRDM1 |
PTPN11 | RB1 | SF3B1 | SLAMF7 | SP140 |
TP53 | TRAF2 | TRAF3 | XBP1 |
Genome-wide copy number variations
500 amplicon panel uniformly covering nearly the entire genome.
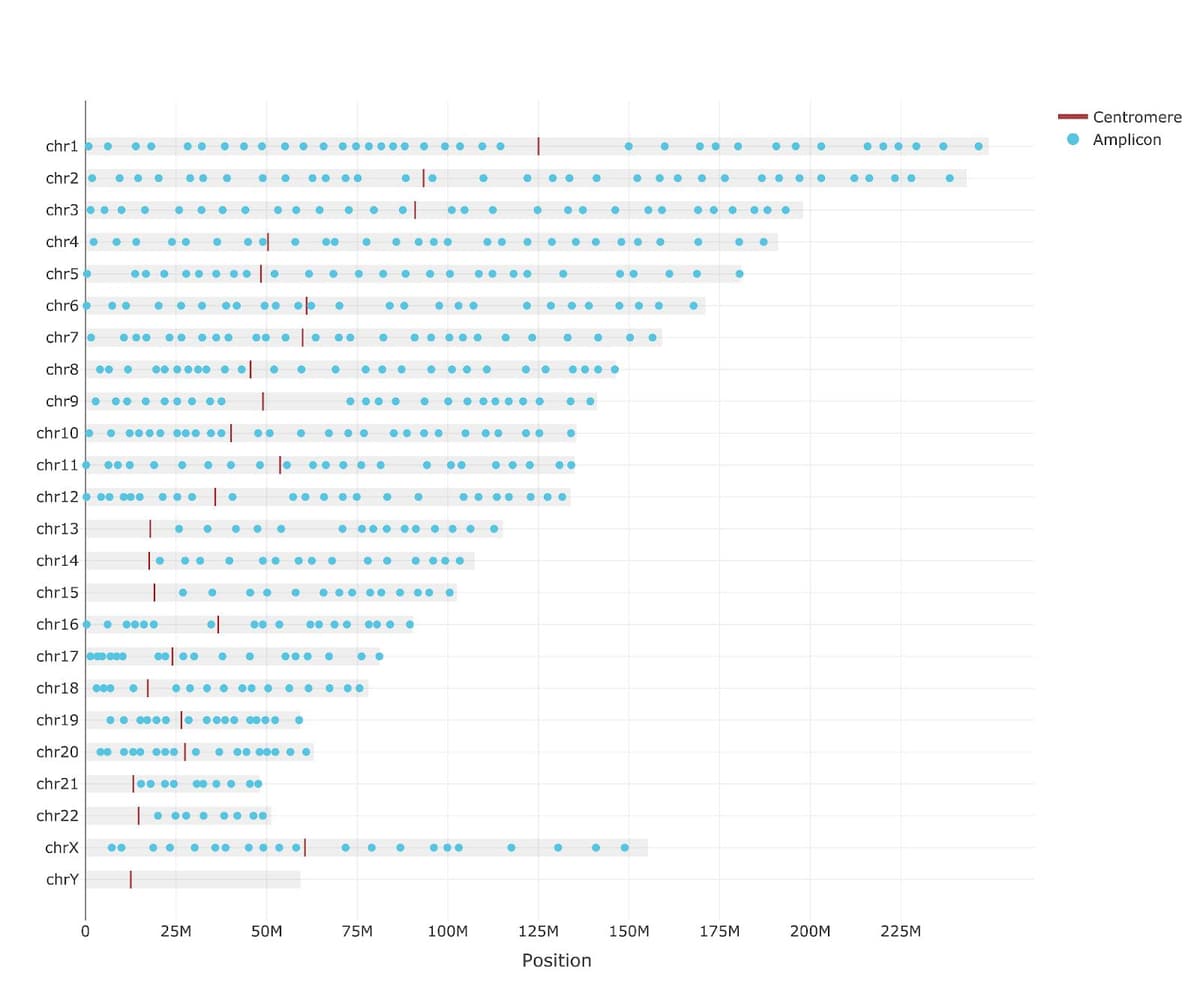
19 antibody oligonucleotide conjugates (AOCs)
Surface antigen makers for cell type classification and therapeutic target identification.

Emerging targets for immunotherapy covered by our assay

The Tapestri Multiple Myeloma Workflow
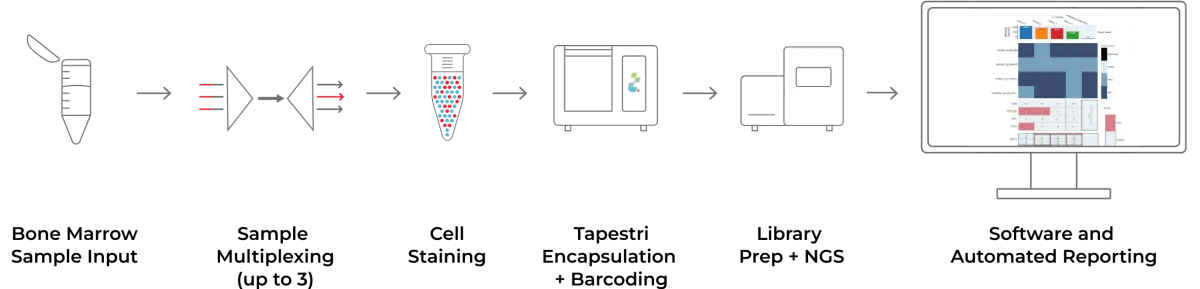
Our streamlined multiple myeloma workflow allows you to multiplex bone marrow samples on the Tapestri Platform, sequence the libraries on an NGS platform, and then easily analyze the results using the Tapestri Multiple Myeloma Software. The Solution can integrate SNVs, focal CNAs, genome-wide CNVs, V(D)J clonotype, and immunophenotype.
Partnering with Mission Bio
To support you and your drug development, we provide full service as a strategic partner for you from single-cell assay design, sample processing, through to data analysis and visualization. Whether we offer services internally or in partnership with qualified CROs, we are dedicated to the accelerated success of your drug development programs.
Featured Resources
Get Started
Contact us to begin your evaluation
For Research Use Only. Not for use in diagnostic procedures.