Why Single Cell for Genome Editing Analysis?
Genome engineering techniques, like CRISPR, result in heterogeneous editing outcomes that include variations in zygosity, off-target effects, co-occurring multiplex edits, chromosomal translocations, and changes to genome stability. Measuring these outcomes accurately cannot be done by traditional sequencing without lengthy clonal outgrowth steps and hours of computational work. With the Tapestri single-cell platform you can obtain critical genome editing measurements to validate your cell therapy or disease model.

Zygosity

Co-Occurrence of Intended Targets

Co-Occurrence of On/Off Targets

Translocations

Immunophenotype

Genome-wide CNV
Create Clear Reports To Easily Interpret Results
Zygosity of On-target Edits

The 10 most frequent variants and the percentage of cells that contain the variant on both alleles (biallelic), one allele (monoallelic), and not at all (wild type, WT). The variant name in the format of “chromosome:position:reference bases/alternative bases” are labeled on the x-axis. For multiplex experiments, the target group may be selected from the dropdown menu in the report.
Co-Occurrence of On-Target Edits

The percentage of cells with the most frequent combinations of multiple on-target edits. Top: frequencies of cells with each combination of edits. Bottom: zygosity of each target.
Genome Editing + Cell Surface Protein

The number of cells with different combinations of edits and cell-surface protein expression. Each column represents cells with the indicated combination of edits and protein expression. Top: the number of cells with each combination. Middle: the zygosity of on-target and predicted off-target edits. Bottom: expression of cell-surface proteins.
Top Translocations

The percentage of cells with the most frequent translocations (up to 10) in the edited sample (# cells shown on the top of each bar). Translocation junctions occur at both predicted on- and off-target editing sites.
Tapestri Genome Editing Solution
Get everything you need to conduct single-cell, multi-omic analysis of gene edits in your lab with this fully-supported, all-in-one package.
- Tapestri Instrument
- DNA Core Kit (8 samples)
- Custom-Designed DNA Panel (+ Optional Protein Panel)
- Software-Based Analysis (via Portal Access or On Premise Activation)
- Optional Antibody Hashing for Sample Multiplexing
- Optional Genome-Wide CNV Panel
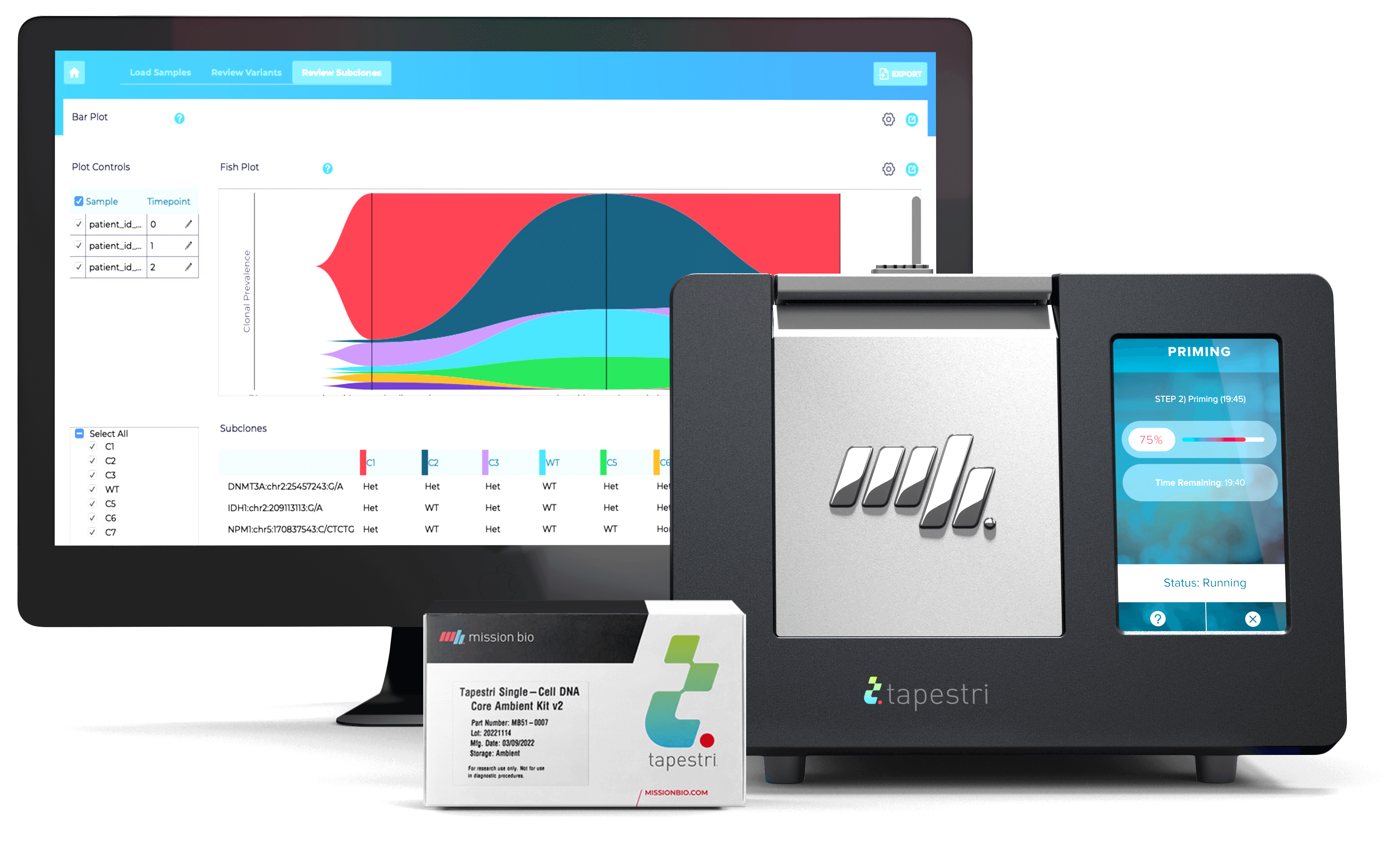
Our streamlined Genome Editing workflow allows you to analyze an edited sample on the Tapestri platform, sequence the libraries on NGS, and then easily analyze the results using the Tapestri Genome Editing Software. The Solution can analyze knockouts (non-homologous end joining) and base edits.
You can generate essential reports with no bioinformatic expertise required. You get clear representations of your data, giving you the insights you need to move forward.
The National Institute of Standards and Technology (NIST) Genome Editing Consortium addresses the measurements and standards needed to increase confidence and lower the risk of utilizing genome editing technologies in research and commercial products. Mission Bio is a proud participating partner.
New Features:
- Addition of whole-genome CNV analysis in Tapestri Genome Editing Software (additional panel required)
- Detect chromosomal translocations – confirm presence of predicted translocation sites
- Multiplex samples with antibody hashing – reduce reagents and labor while increasing sample throughput
The Tapestri Genome Editing Workflow
The end-to-end Tapestri workflow for single-cell genome editing analysis. A sample of gene-edited cells are processed on the Tapestri Platform, sequenced using NGS, and analyzed using Tapestri Genome Editing software. The assessment at single-cell resolution of on-targets, predicted off-targets, zygosity of edits, co-occurring (multiplex) edits, predicted translocations, cell-surface proteins, and genome integrity is possible. The Solution can be used to analyze gene knockouts (via non- homologous end joining) and base edits.
- Step 1 – Design Panel & Run Edited Samples on Tapestri®
- Step 2 – Sequence Libraries on NGS
- Step 3 – Software Runs Automated Analysis and Generates Intuitive Data Report
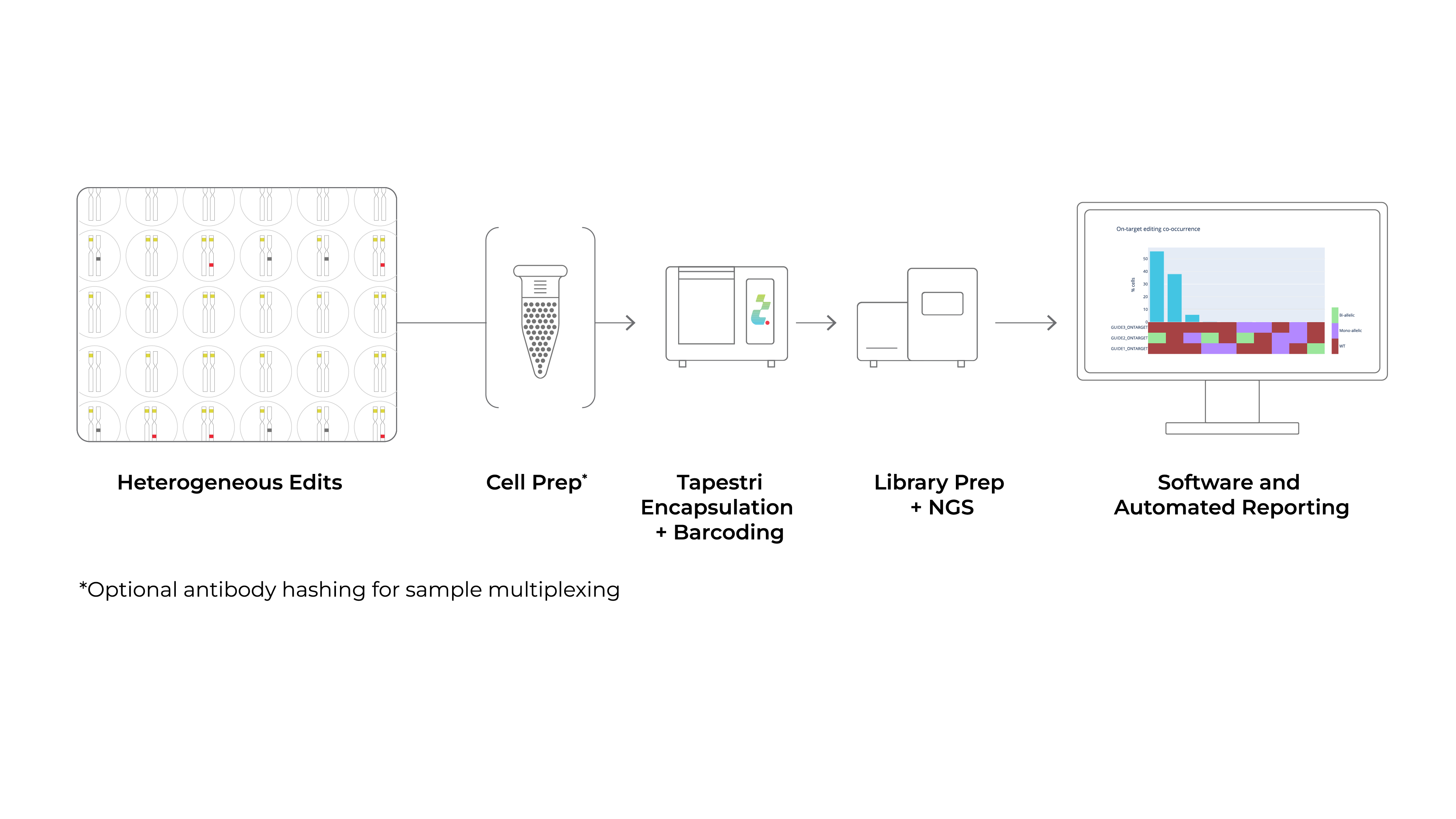
Effortlessly analyze on- and off-target edits, assess zygosity, predicted translocations, co-occurring edits, and genome-wide copy number variations. Our automated analysis and intuitive reports make the process easy and efficient.
Primary Applications

"This multi-omic genome-editing solution will enable critical insights for gene-edited cell therapies beyond what conventional bulk analysis can offer. The automated report gives an immediate and intuitive first look at our data, something that would usually require hours of bioinformatic labor. All in all, we are better equipped to understand our editing results with this technology."
Saar Gill MD, PhD, Associate Professor of Medicine at the University of Pennsylvania
Get Started
Contact us to discuss your project:
Fill out the form below and somebody from our team will get back to you within 2 business days.