Why Single Cell?
Over the past decade, numerous advances in biotechnology have translated to improvements in the medical arena. Yet, limitations in analytical tools still remain and present significant roadblocks to the study of pathologies and the development of life-changing therapies.
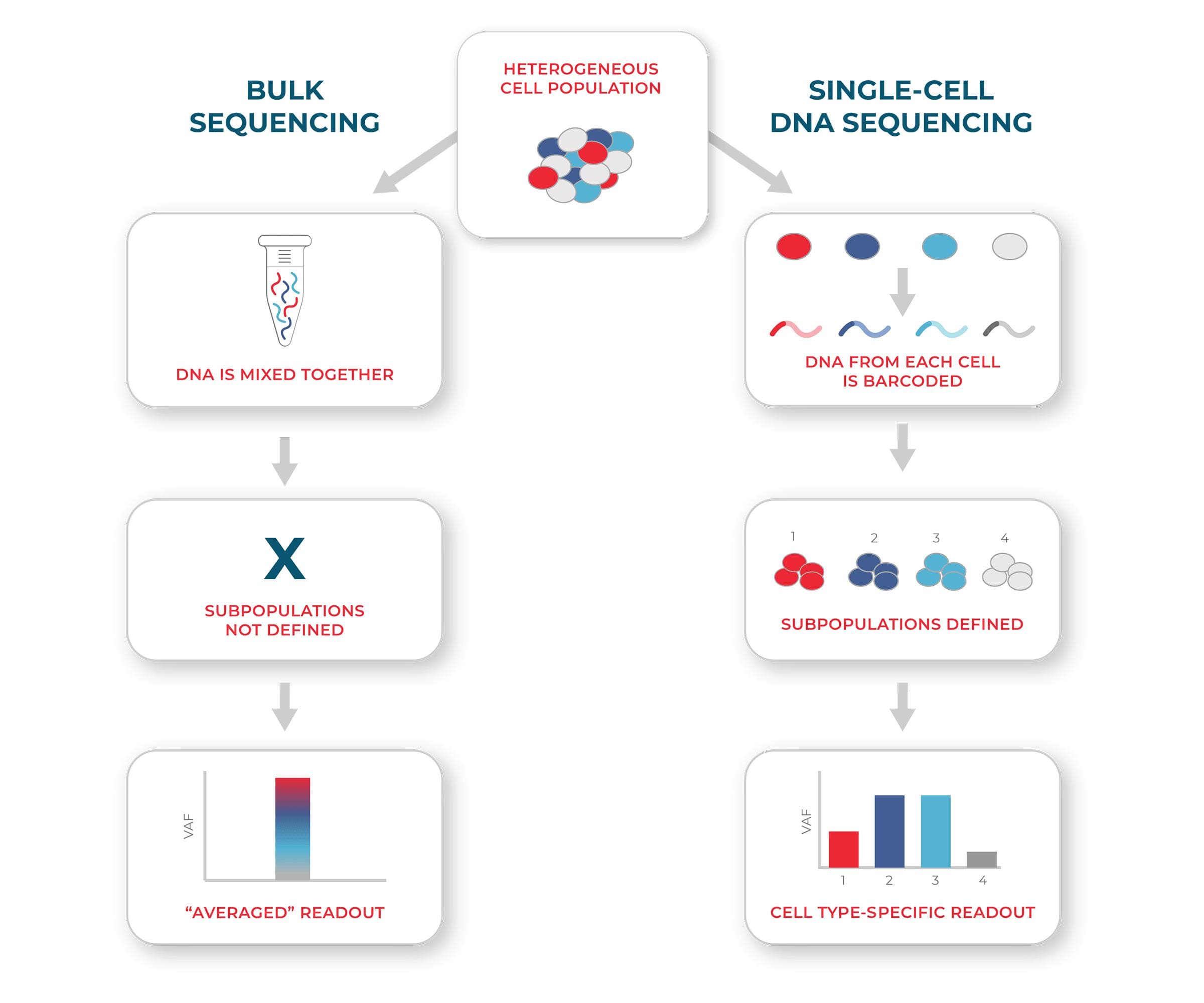
Today, many genotyping workflows rely on bulk assays, which mix DNA across cells and report average readouts. Although appropriate for some questions, these assays do not preserve the information of individual cells, and therefore provide only limited knowledge about biological complexity. Single-cell DNA sequencing, on the other hand, evaluates the genotype of individual cells, and thus provides a richer picture of the sample.
Single-cell sequencing allows for direct measurement of mutations in individual cells by first encapsulating the cells in small droplets and then introducing individual molecular barcodes, enabling researchers to confidently identify which mutations are present in which cells, and no longer need to infer things like zygosity and mutational co-occurrence.
This is especially pertinent for applications where information at the cellular level matters. Some of the most pressing challenges include elucidating the complexity of heterogeneous diseases like cancer, and the underlying mechanisms driving them. Other hurdles lie within the process of pharmacological development, in which each therapeutic candidate must be characterized with sufficient granularity to ensure safety and efficacy.
Single cell Sequencing vs Bulk Sequencing
Why Multi-omics?
Conventional workflows in disease research and therapeutic development involve running multiple separate assays, often on different instruments, to get answers. This approach has several drawbacks.
First, precious samples that are difficult to obtain must be used in multiple assays — leaving little for downstream analyses. Second, the data integration and analysis of multi-assay studies is complex and typically requires bioinformatics expertise to glean correlations across different datasets. Third, conducting multiple assays on different “omics” platforms can get expensive, and will often require longer timelines before researchers get answers to their experimental questions.
Tapestri’s Multi-omics Solution
Fortunately, innovative multi-omics technologies like the Tapestri Platform are offering a solution to these issues. Tapestri simultaneously measures DNA and cell-surface proteins across thousands of individual cells.
By collapsing genotypic and phenotypic assays into a single workflow, Tapestri both preserves precious samples and enables multi-parameter data to be gathered quickly and easily. This powerful approach is offering an unprecedented look into the nuances of biology and transforming how we explore disease and develop therapies.
The Impact of Single-Cell and Multi-Omics Technologies
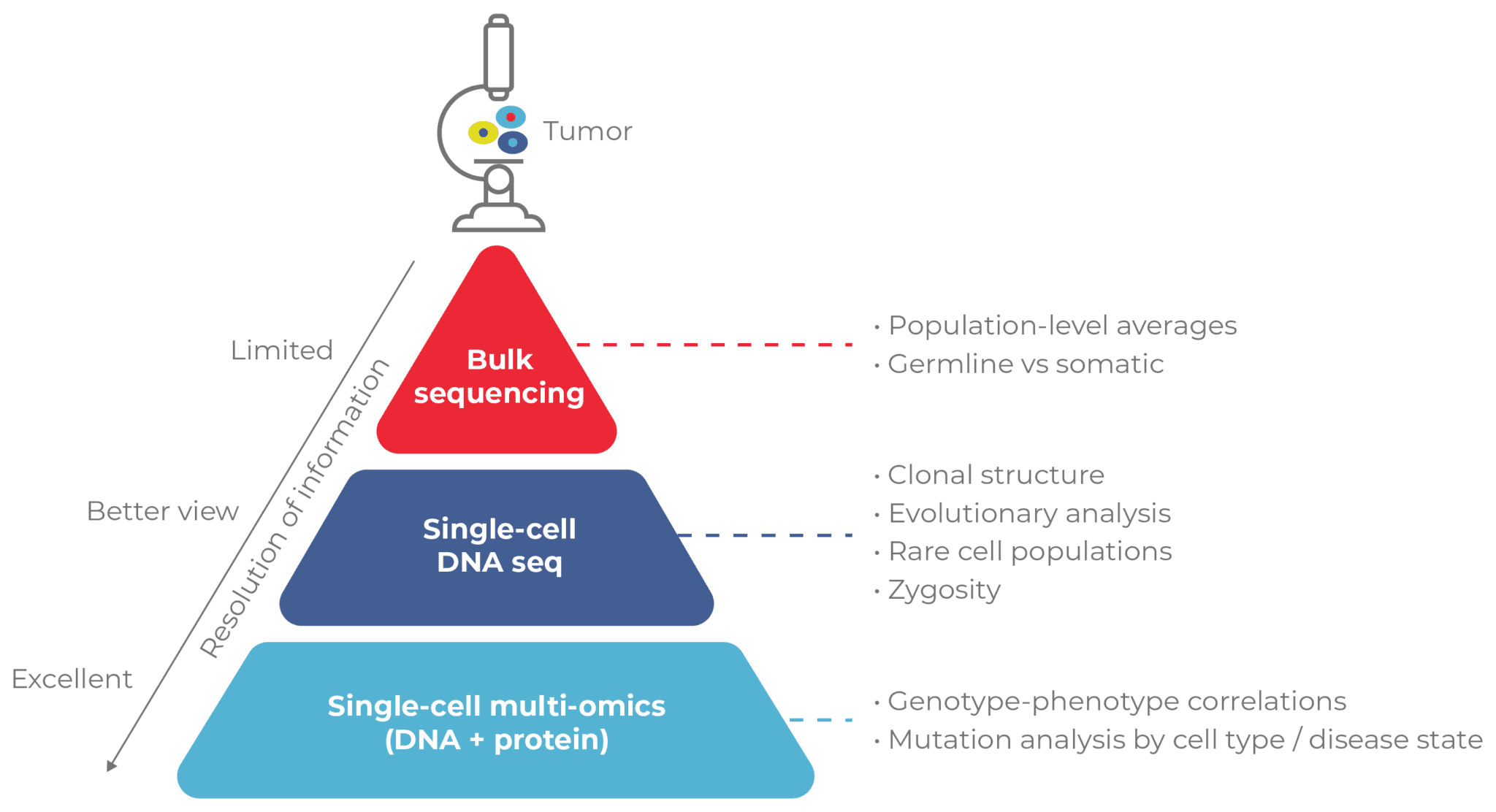
Single-cell Applications
Single-cell analysis is advancing several scientific areas, including oncology and the development of advanced therapeutics. For instance, Tapestri is helping advance cancer translational research, where information about tumor architecture can be used to inform diagnosis, prognosis, and treatment strategies in a highly personalized fashion.
Tapestri is also helping advance the development of novel cell and gene therapies. By measuring multiple genotypic alterations across cells, therapy developers achieve deeper characterization of gene-modified cell therapies and assess in vivo gene therapies in model systems.
The benefits of single-cell analysis extend to academia and industries alike. Pharma and biopharma can capitalize on the highly sensitive analysis of Tapestri to assess therapy resistance, MRD, and characterize cell-based drug candidates.
Ready to learn more?
Use Tapestri Designer to design a custom panel tailored to your needs.