Poster
Method to analyze mutational and phenotypic profiles from single cell for clonal evolution
This poster demonstrates the capability of the Tapestri Platform multi-omics workflow (which analyzes the DNA and protein information from the same cells) to discriminate cell populations. Using a two-donor PBMC mixture as a model system, we show how processing on Tapestri identifies two clones using the genetic variants and multiple cell types including T-cells, B-cells, monocytes, and NK-cells using phenotypic expression. Moreover, Tapestri identified the proportion of each cell type belonging to each of the two donors.
SHARE THIS PAGE
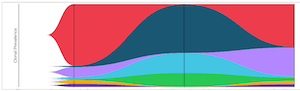
Single-cell multiomic clonal tracking in myeloma identifies SMM clones that progress to MM and low frequency MM clones with resistance features enabling more precise application of targeted therapies
Adam Sciambi
EHA (2024)

Single-Cell MRD Assessment in AML Reveals Clonal Diversity and Genotype–Phenotype Discordance Missed by Bulk Methods
Adam Sciambi, Daniel Mendoza, Kathryn Thompson, Lan W. Beppu, Benjamin Geller, Indira Krishnan, Lubna Nousheen, Shu Wang, Charlie Murphy, Jerald P. Radich, and Todd E. Druley
(2025)

Comprehensive On-and Off-target Confirmation Using Integrated rhAmpSeq and Targeted DNA Sequencing Single-Cell Technology
Indira Krishnan, Shu Wang, Saurabh Gulati
ASGCT 2025 (2025)

High Throughput Single-cell Assessment of Genome Integrity and Toxicity Events Associated With Edited Cells
Chieh-Yuan (Alex) Li; Saurabh Parikh; Saurabh Gulati; Donjo Ban; Nechama Kalter; Michael Rosenburg; Qawer Ayaz; Joanne Nguyen; Benjamin Miltz; Yang Li1; Madhumita Shrikhande; Edward Szekeres; Ayal Hendel; Benjamin Schroeder; Shu Wang
ESGCT 2024 (2024)